Knowledge Graph
The dataset is parsed into a structured graph of 50 drugs, 41 conditions, and 68 side effects. These are the indexes that power the retrieval engine.
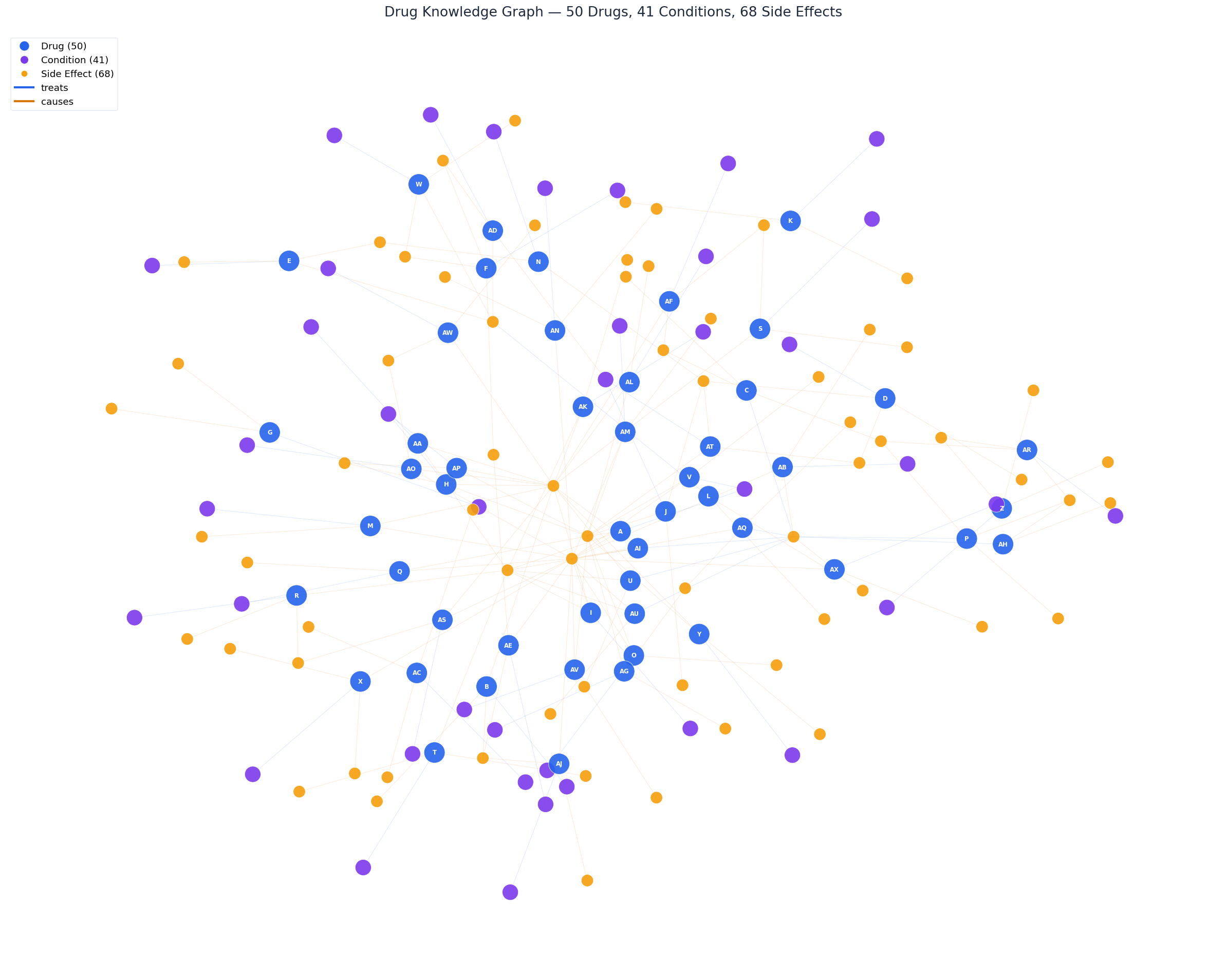
All 50 drugs (blue), 41 medical conditions (purple), and 68 side effects (yellow) connected by "treats" and "causes" relationships. This is the graph that our 3-layer retrieval engine queries at runtime.
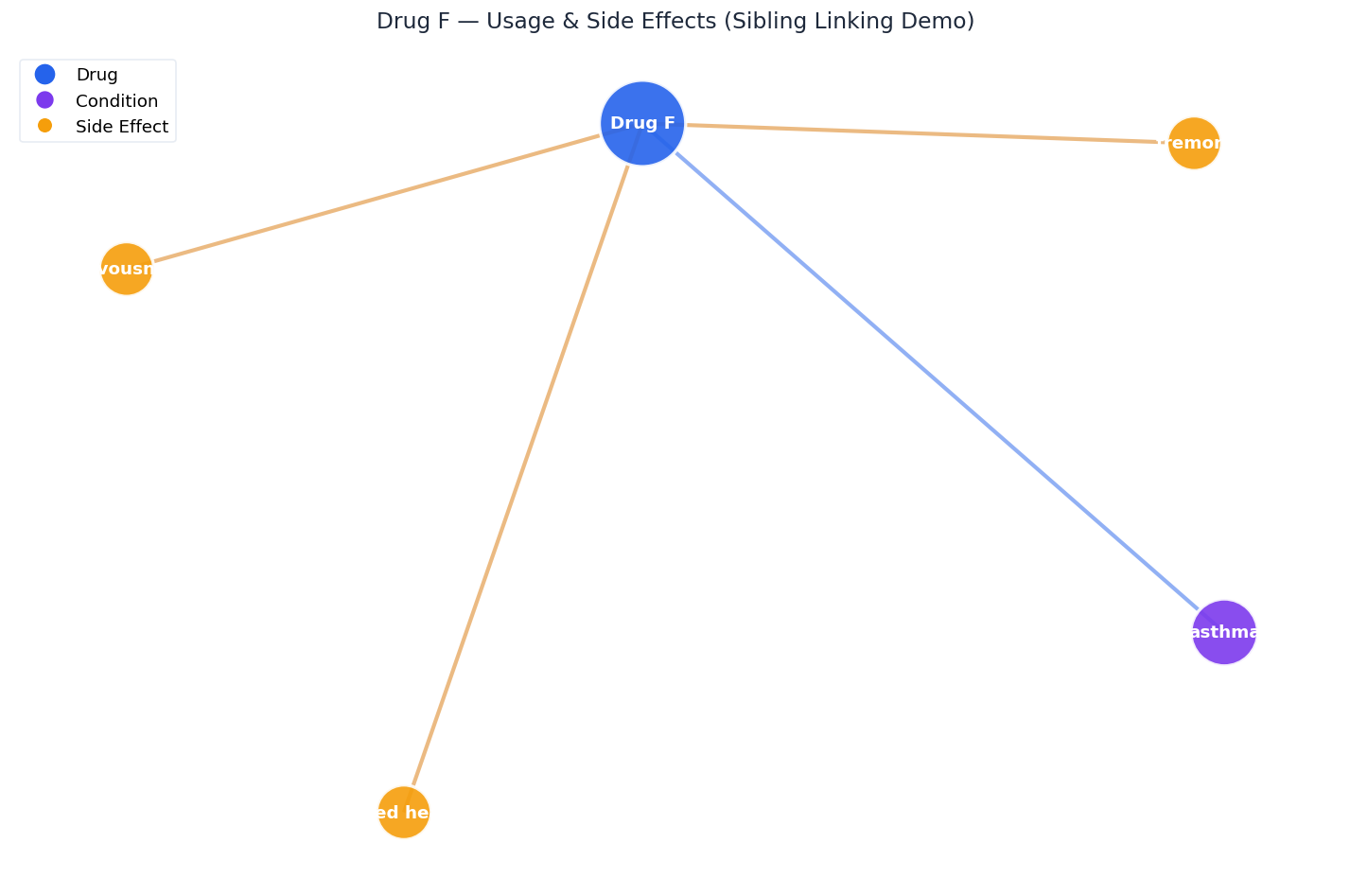
Drug F treats asthma (purple) and causes tremors, nervousness, and increased heart rate (yellow). The sibling linker ensures both the usage doc and side-effect doc are always retrieved together -- this is why multi-hop queries work.
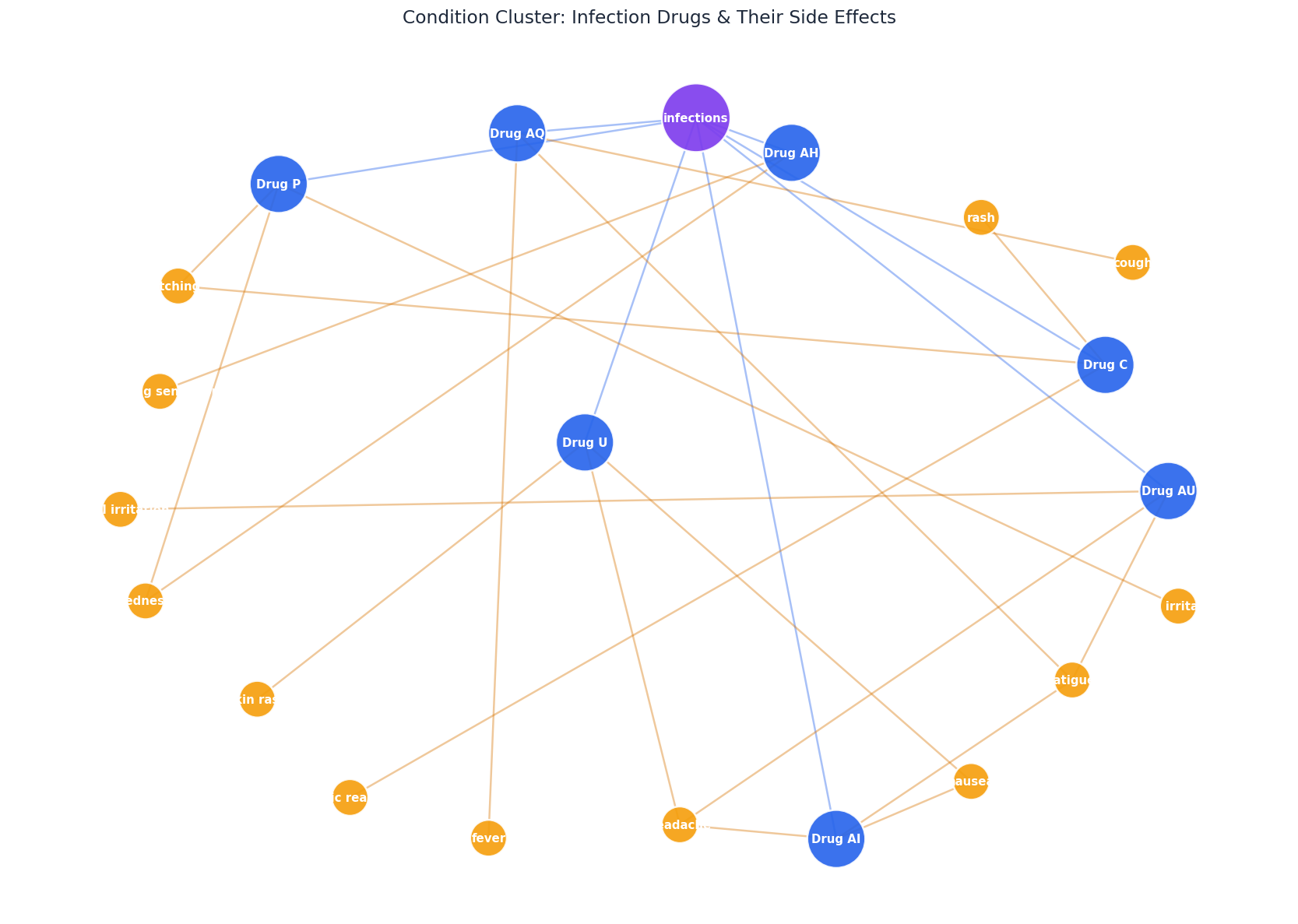
All drugs that treat infections, clustered around the shared condition node. This is what the retrieval engine sees when you ask "Which drugs are used for infections?" -- the condition index maps directly to this subgraph.